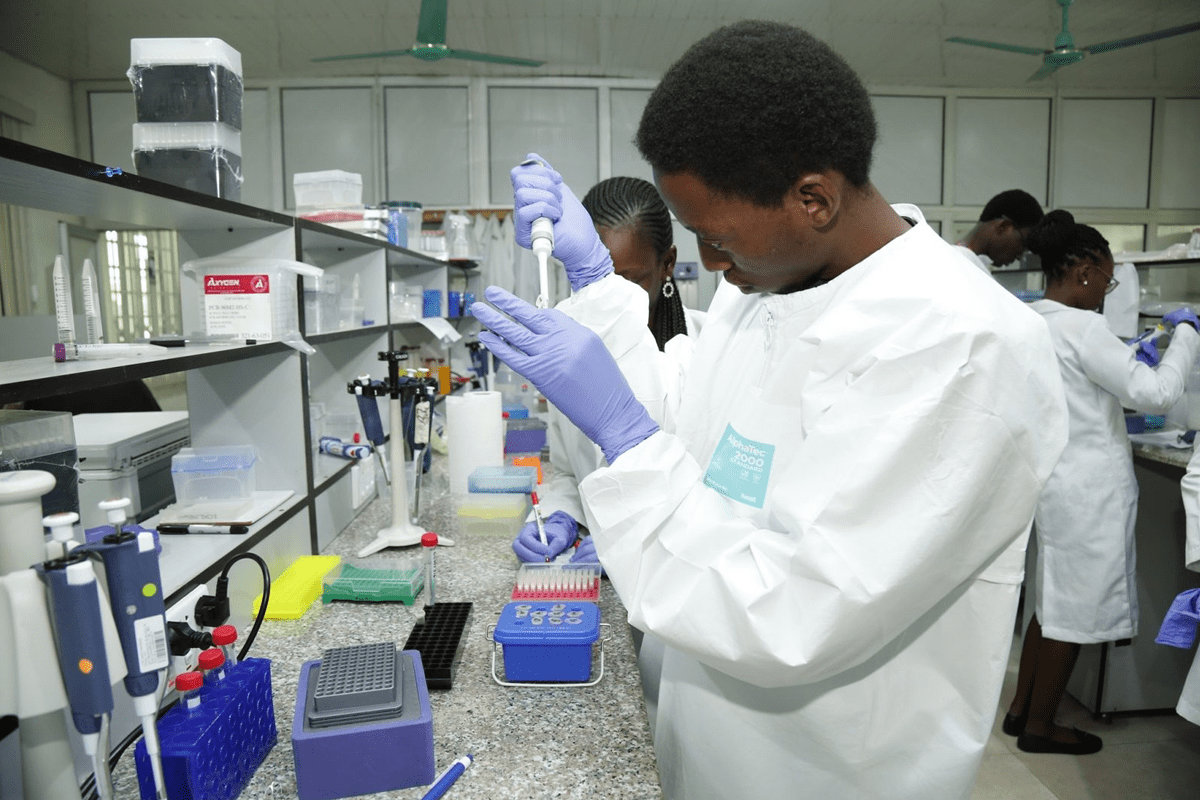
EDE, NIGERIA, August 17, 2021 The African Centre of Excellence for Genomics of Infectious Diseases (ACEGID) has started an intensive hands-on bioinformatics workshop on the analysis of SARS-CoV-2 genomic and...
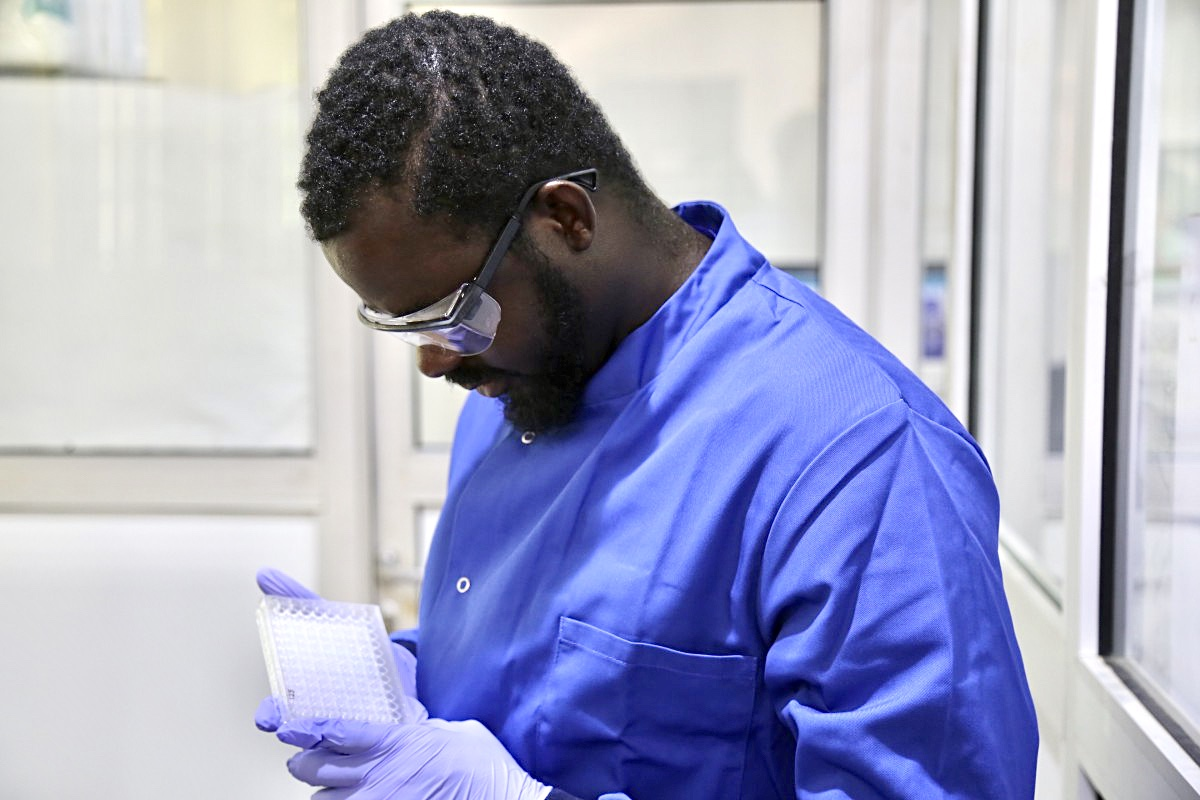

Globally, infectious diseases account for the largest burden of diseases of public health importance. They also remain a never-ending challenge as new pathogens causing severe infectious diseases are constantly emerging...
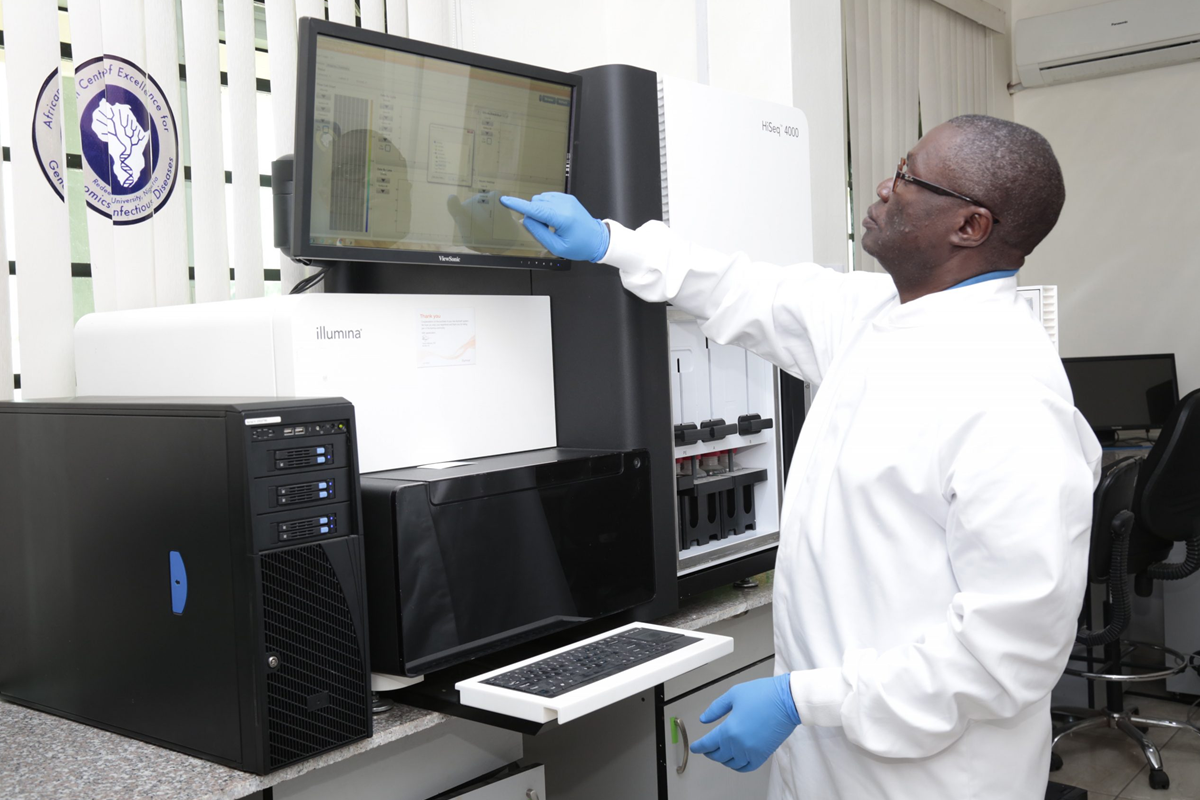

ACEGID's approach, results and impacts were the focus of a recent World Bank Results Brief. The publication highlights how we are countering immediate public health threats as well as detecting...

Towards scaling up genomic sequencing capacity on the African continent, the African Centre of Excellence for Genomics of Infectious Diseases (ACEGID), in partnership with the Africa Pathogen Genomics Initiative, is
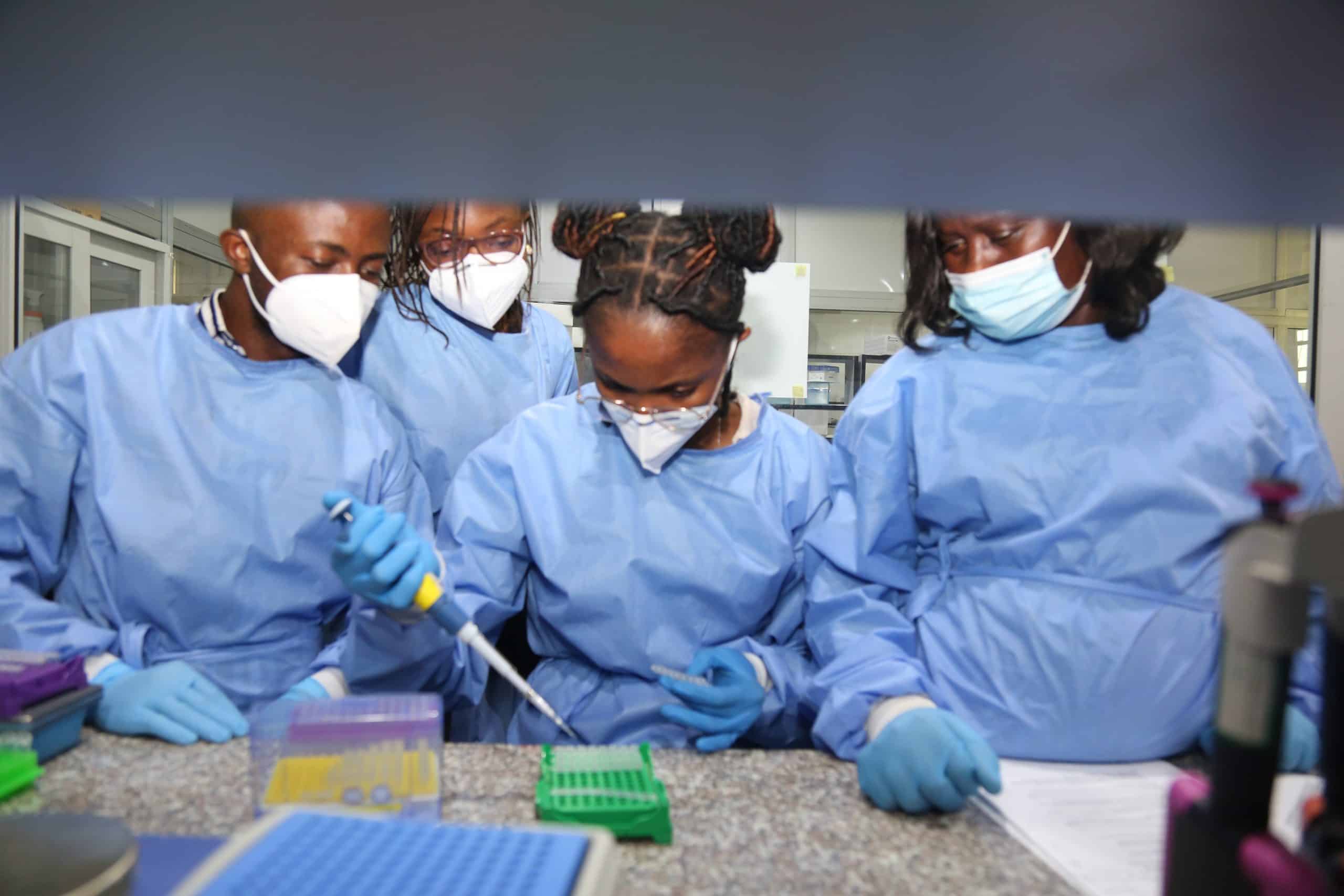

ACEGID and Illumina to Increase Africa’s Genomics Capacity through Establishment of a Joint Training Academy
Today, the African Centre of Excellence for Genomics of Infectious Diseases (ACEGID), Redeemer’s University, announced that it will collaborate with Illumina, a global leader in sequencing and array-based technologies, on a joint workforce development training academy, with the objective of increasing Africa’s genomics capacity.
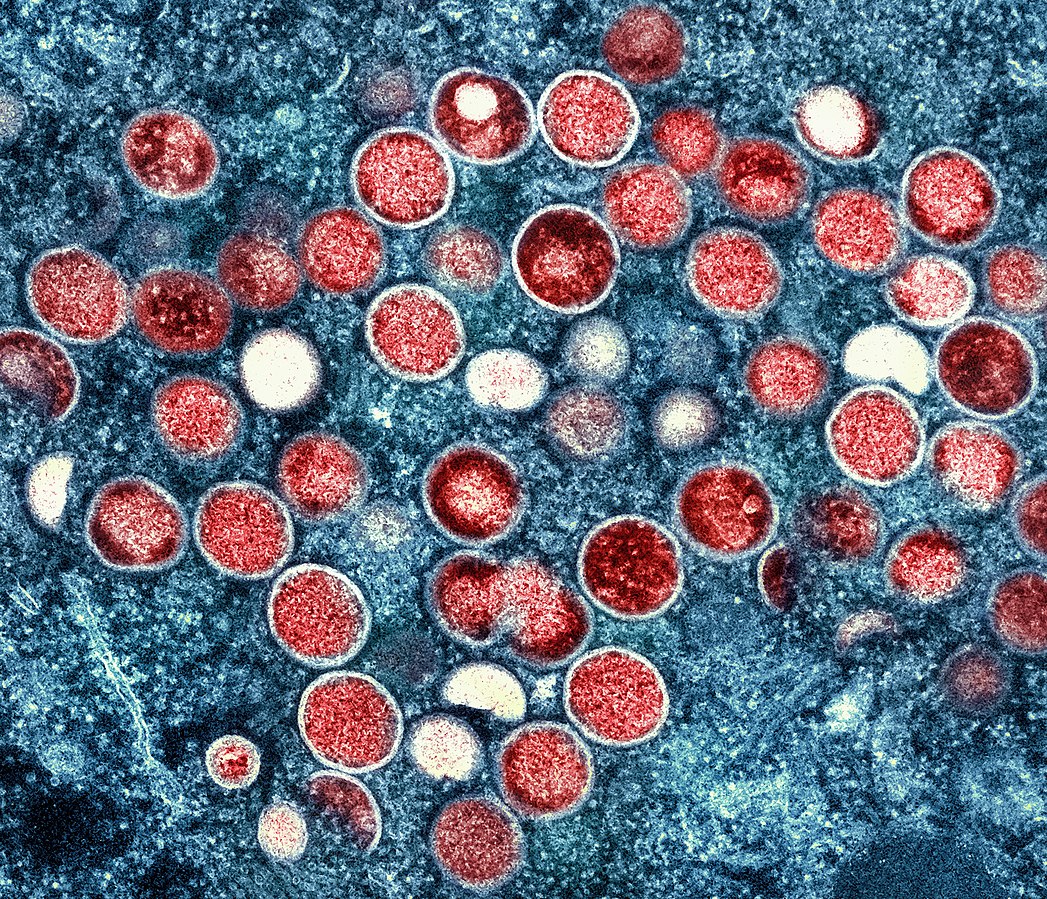
Monkeypox Pandemic reflects Global Neglect of Africa
Since the recent announcements of monkeypox outbreak in Europe and America, Prof. Christian Happi has granted several interviews to journalists across the world. Here, we present some of the recurring themes of such conversations. These are highlights of our position on the ongoing monkeypox pandemic.
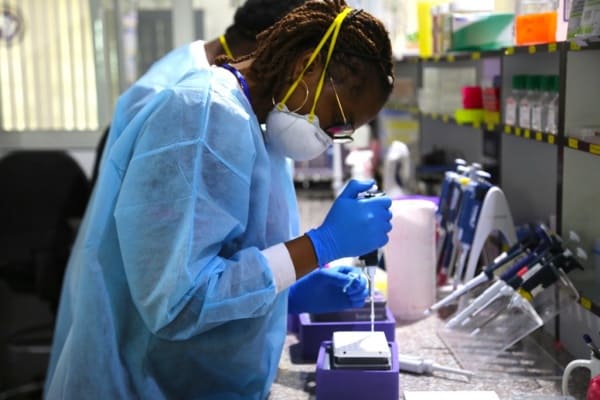

African Center of Excellence for Genomics of Infectious Diseases
ACEGID,Nigeria
Our Mission

Our Partners






