ACEGID facilitated a three-week training on SARS-CoV-2 sequencing and bioinformatics for Rwandan scientists at the Rwanda Biomedical Centre (RBC).
The training was held as a collaboration between ACEGID and RBC from January 31 to February 18, 2022.
ACEGID scientists trained participants in the principles and practices of SARS-CoV-2 sequencing steps including sample handling, extraction, amplification, quantification, library preparation and sequencing. The participants had hands-on training on Illumina sequencing with Miseq and Oxford Nanopore Technology (ONT) MinION.
Participants were also trained in bioinformatics. They learnt how to assemble sequences, how to submit data to online repositories, how to identify variants and how to interpret data.
Twenty-two scientists from three Rwandan public health institutions were trained in the workshop. 122 SARS-CoV-2 genome sequences (50 on ONT MinION and 72 on Illumina Miseq) were assembled during the training.























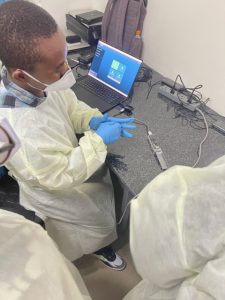
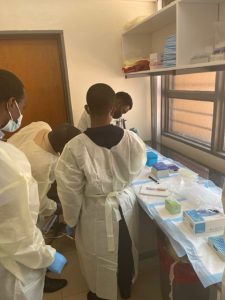
Participants’ feedback showed enthusiasm for applying the lessons learnt towards fighting infectious diseases in Rwanda. Below are excerpts:
